spesim recipes: interspecific interactions
Source:vignettes/spesim-recipes-interactions.Rmd
spesim-recipes-interactions.RmdThis recipe shows how to specify simple neighbour
effects using INTERACTIONS_EDGELIST.
Values:
-
< 1suppressive (competition) -
> 1facilitative -
= 1neutral
Baseline without interactions
library(spesim)
#> spesim 0.5.2 loaded - try spesim_run() to generate a simulation.
P <- load_config(system.file("examples/spesim_init_basic.txt", package = "spesim"))
#> ========== INITIALISING SPATIAL SAMPLING SIMULATION ==========
P$N_SPECIES <- 10
P$N_INDIVIDUALS <- 2000
P$ADVANCED_ANALYSIS <- FALSE
# Turn interactions off
P$INTERACTION_RADIUS <- 0
P$INTERACTIONS_EDGELIST <- NULL
set.seed(P$SEED)
res0 <- spesim_run(P, write_outputs = FALSE, seed = P$SEED, interactions_print = FALSE)
#> spesim: running simulation
#> Note: no non-1 interactions for species: A, B, C, D, E, F, G, H, I, J.
plot_spatial_sampling(res0$domain, res0$species_dist, res0$quadrats, res0$P)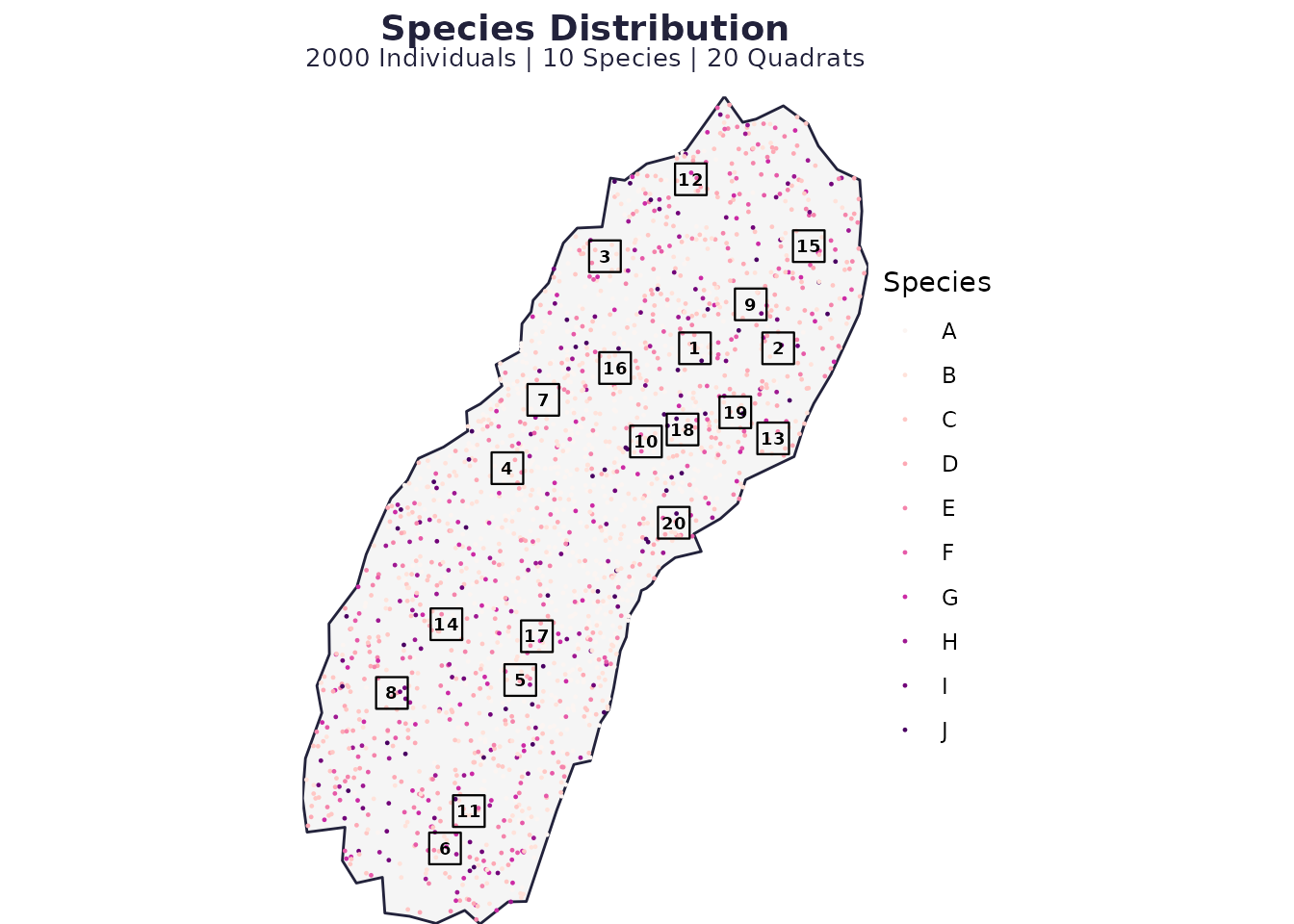
Add a few directed interaction rules
Here, species A suppresses B..D locally, and species C facilitates A.
P2 <- P
P2$INTERACTION_RADIUS <- 2
P2$INTERACTIONS_EDGELIST <- c(
"A,B-D,0.7",
"C,A,1.3"
)
res1 <- spesim_run(P2, write_outputs = FALSE, seed = P2$SEED, interactions_print = TRUE)
#> spesim: running simulation
#> Note: no non-1 interactions for species: E, F, G, H, I, J.
#> ---- Interactions Summary ----
#> Species: 10 (A..J)
#> Radius : 2
#> Non-1 entries: 4 (4.0% of 100)
#> Asymmetry (mean |M - t(M)|): 0.024
#>
#> Matrix ('.' = 1):
#> A B C D E F G H I J
#> A . 0.7 0.7 0.7 . . . . . .
#> B . . . . . . . . . .
#> C 1.3 . . . . . . . . .
#> D . . . . . . . . . .
#> E . . . . . . . . . .
#> F . . . . . . . . . .
#> G . . . . . . . . . .
#> H . . . . . . . . . .
#> I . . . . . . . . . .
#> J . . . . . . . . . .
#>
#> Non-1 entries (sorted by |value-1|):
#> focal neighbor value
#> A B 0.7
#> A C 0.7
#> A D 0.7
#> C A 1.3
#> ------------------------------
# Visualise the outcome
plot_spatial_sampling(res1$domain, res1$species_dist, res1$quadrats, res1$P)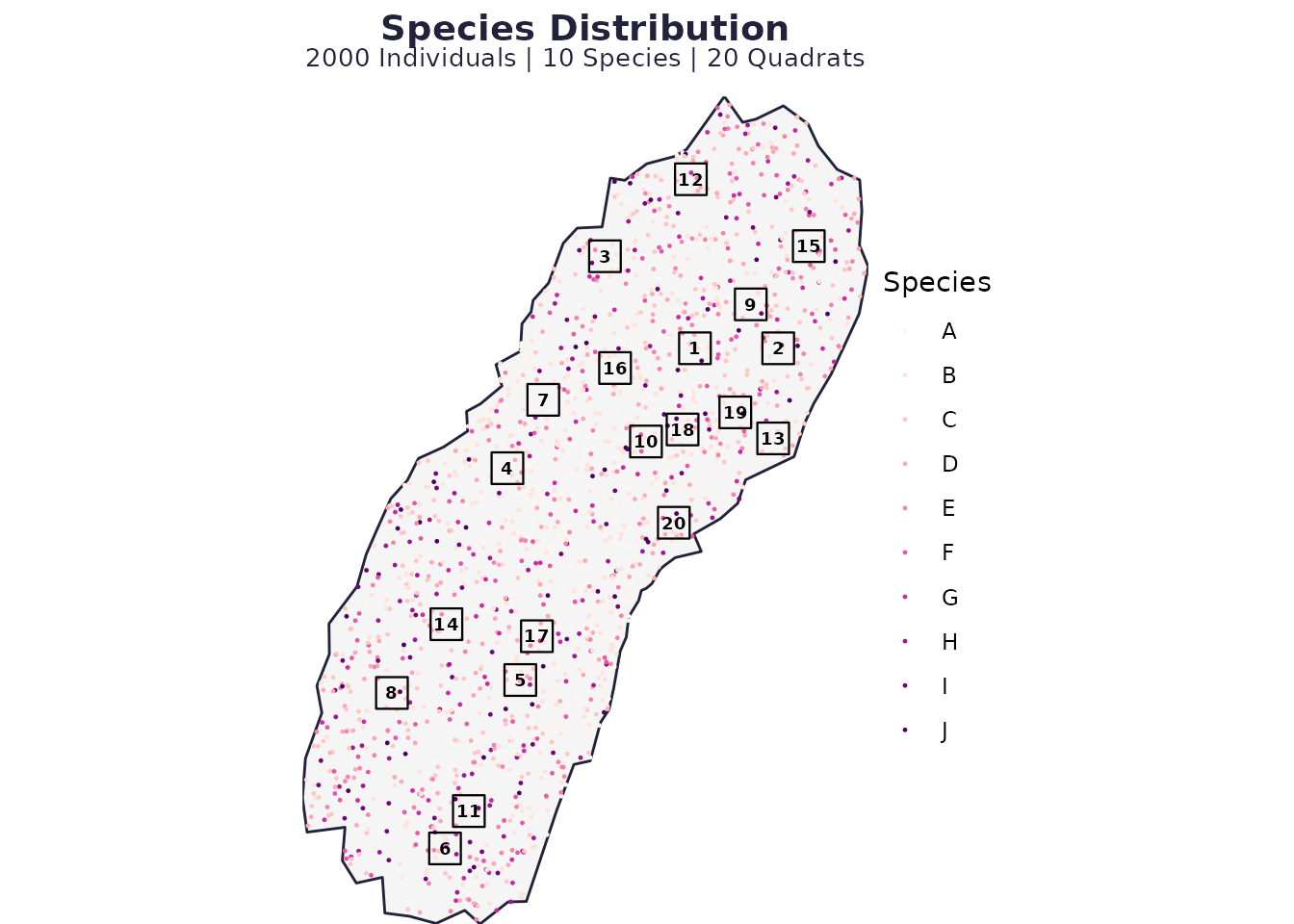
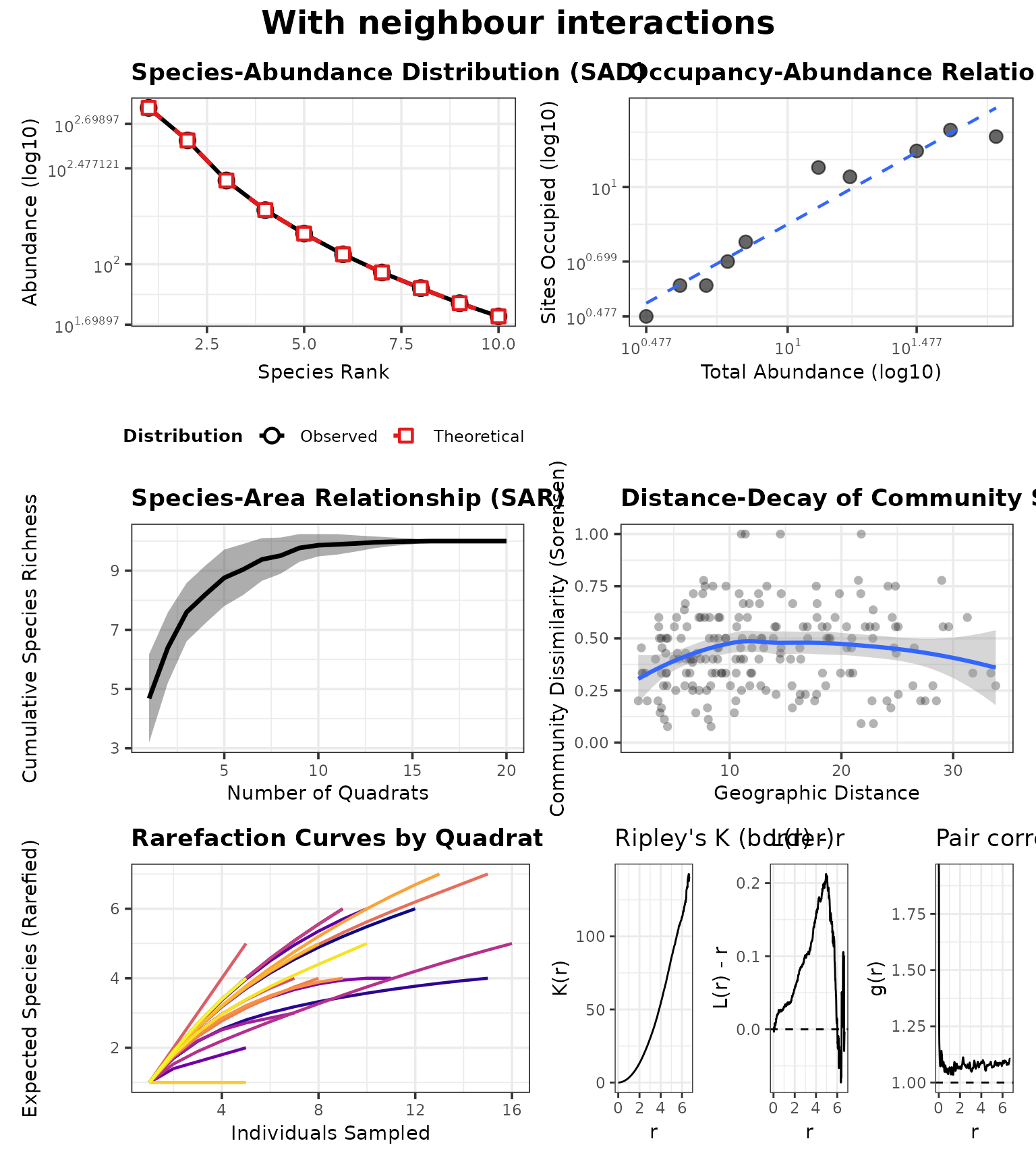
Consequences in the advanced analysis panel
Neighbour effects can change co-occurrence patterns and local abundance, which in turn changes what you observe in quadrats.
In ecological terms, competition (coefficients < 1) can create spatial segregation and reduce local richness in neighbourhoods dominated by a strong competitor, while facilitation (> 1) can increase local co-occurrence.
In the advanced panel, interaction effects can manifest as:
- shifts in occupancy–abundance (suppressed species become rarer and occupy fewer sites),
- changes in distance–decay (if interactions amplify patchiness),
- and altered rarefaction curves (effective evenness/richness per site).
panel0 <- generate_advanced_panel(res0) + patchwork::plot_annotation(title = "Baseline: no interactions")
panel1 <- generate_advanced_panel(res1) + patchwork::plot_annotation(title = "With neighbour interactions")
panel0
#> `geom_smooth()` using formula = 'y ~ x'
#> `geom_smooth()` using formula = 'y ~ x'
#> Warning: Removed 1 row containing missing values or values outside the scale range
#> (`geom_line()`).
#> Removed 1 row containing missing values or values outside the scale range
#> (`geom_line()`).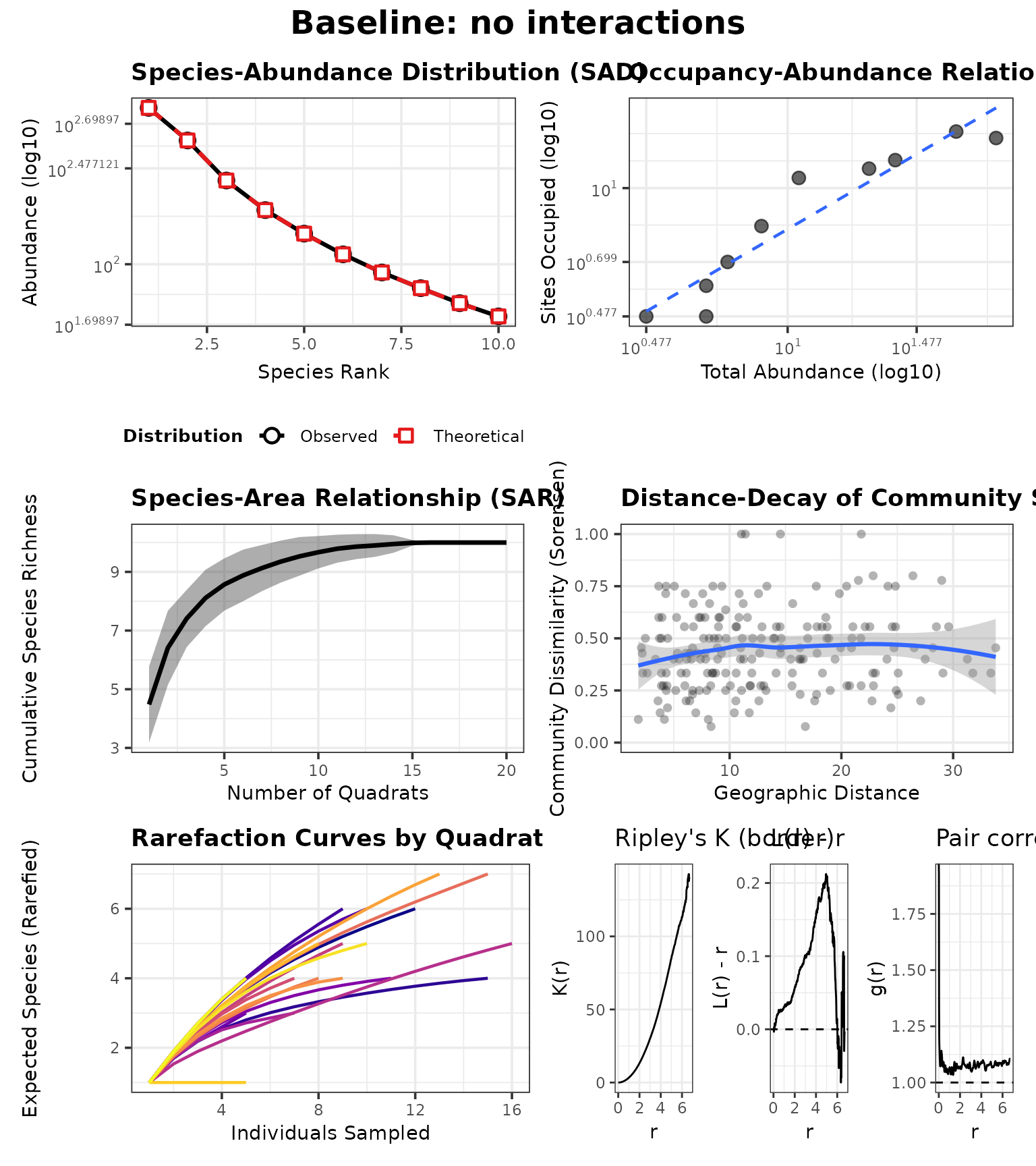
panel1
#> `geom_smooth()` using formula = 'y ~ x'
#> `geom_smooth()` using formula = 'y ~ x'
#> Warning: Removed 1 row containing missing values or values outside the scale range
#> (`geom_line()`).
#> Removed 1 row containing missing values or values outside the scale range
#> (`geom_line()`).