Quadrat placement schemes in spesim
Source:vignettes/spesim-quadrat-placement.Rmd
spesim-quadrat-placement.RmdThis vignette shows the five quadrat placement schemes available in spesim, with reproducible code and simple visualisations for each:
-
random→ [place_quadrats()] -
tiled→ [place_quadrats_tiled()] -
systematic→ [place_quadrats_systematic()] -
transect→ [place_quadrats_transect()] -
voronoi→ [place_quadrats_voronoi()]
All schemes place axis-aligned rectangular quadrats (i.e., not rotated). Each function attempts to ensure quadrats lie fully within the sampling domain; specific constraints differ by scheme.
Setup: a shared irregular domain
library(spesim)
library(sf)
library(ggplot2)
set.seed(1)
domain <- create_sampling_domain()
# Quadrat size is given in the domain's coordinate units.
# The default domain is synthetic and uses an arbitrary planar scale.
quadrat_size <- c(1.5, 1.5)A small helper to plot the domain + quadrats consistently:
# Plotting helper (exported by spesim)
# See ?plot_quadrats for arguments.1) Random (non-overlapping) placement
Use when: you want simple random sampling and you don’t want quadrats overlapping.
Mechanics: rejection sampling of random centres (uniform over a shrunk bounding box), accepting only quadrats that are fully inside the domain and do not overlap already accepted quadrats.
set.seed(2)
q_random <- place_quadrats(domain, n_quadrats = 20, quadrat_size = quadrat_size)
plot_quadrats(domain, q_random, "Random placement (non-overlapping)")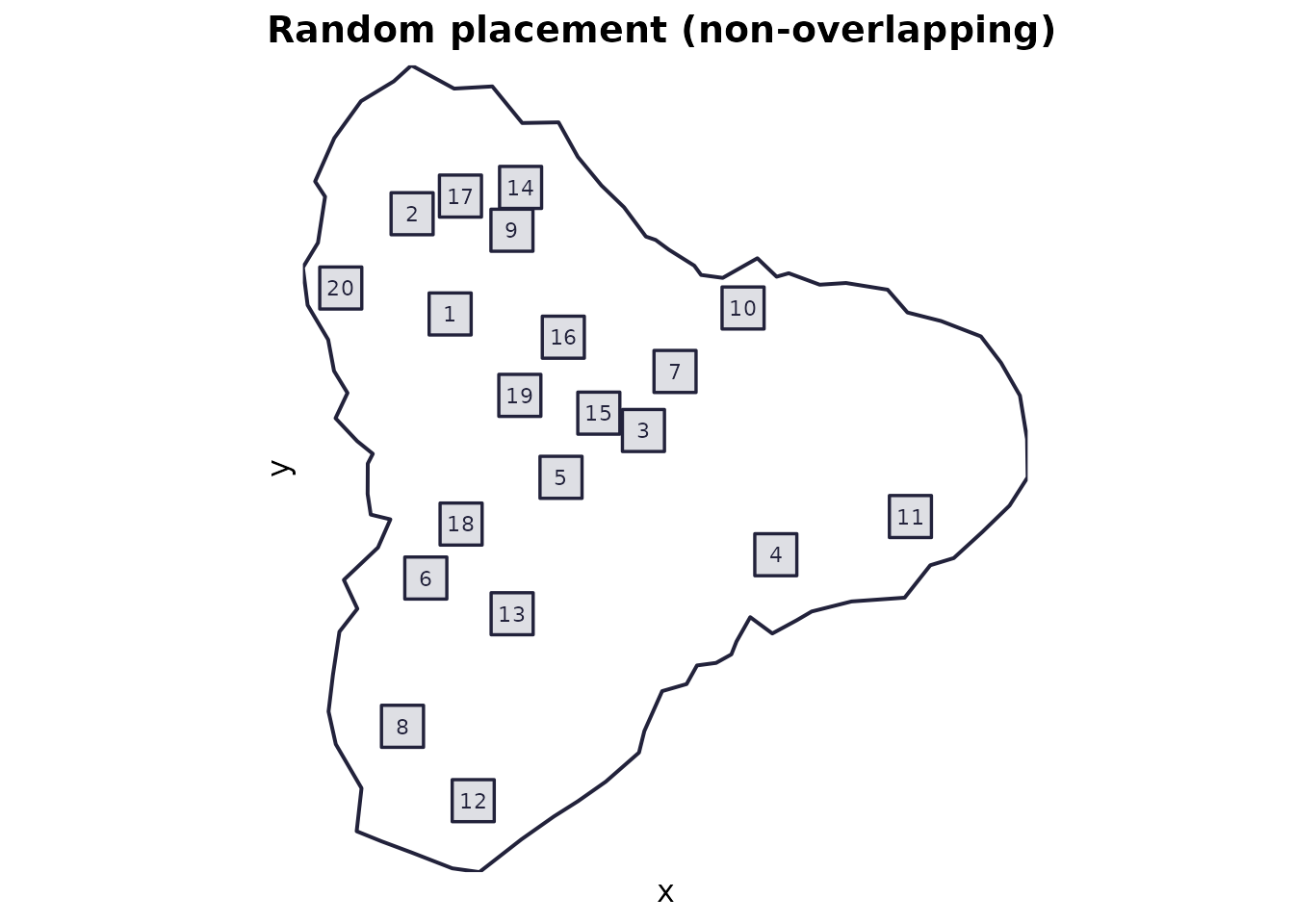
Notes:
- If the domain is small or the quadrats are large, you may get fewer quadrats than requested (with a warning).
- This is often a good default for teaching.
2) Tiled placement
Use when: you want quadrats drawn from a regular tiling, with strict non-overlap.
Mechanics: a regular grid of cells
(st_make_grid) at the quadrat size is created; tiles fully
inside the domain are candidates; n_quadrats are sampled
from these candidates.
set.seed(3)
q_tiled <- place_quadrats_tiled(domain, n_quadrats = 20, quadrat_size = quadrat_size)
plot_quadrats(domain, q_tiled, "Tiled placement (grid cells inside domain)")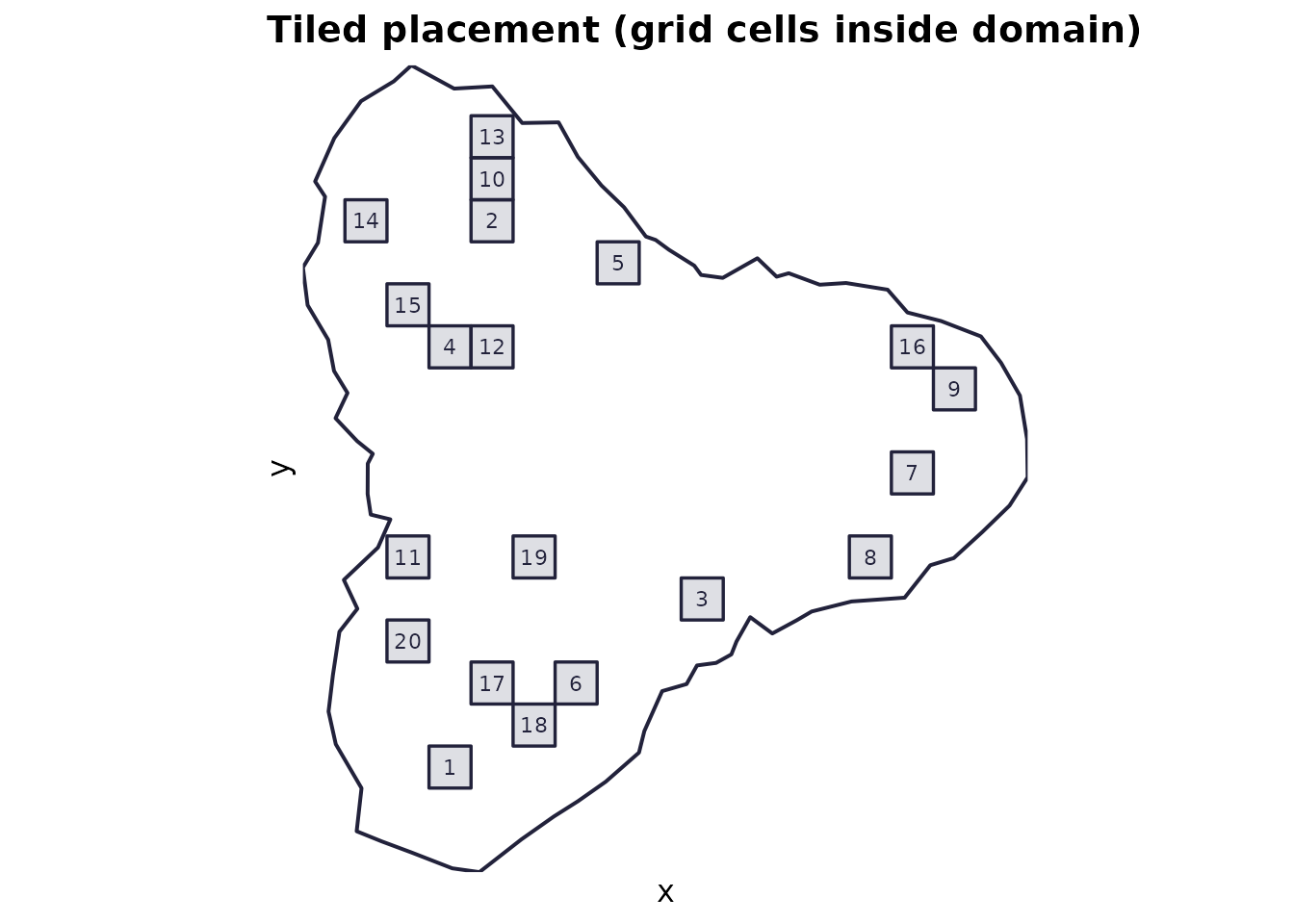
Notes:
- Returned quadrats never overlap.
- If the domain has narrow “necks”, many grid cells may be excluded.
3) Systematic placement
Use when: you want a regular lattice of quadrats (systematic sampling).
Mechanics: centres are placed on an
nx × ny grid spanning the domain bounding box; quadrats
centred on those points are retained only if they fit entirely in the
domain.
set.seed(4)
q_systematic <- place_quadrats_systematic(domain, n_quadrats = 25, quadrat_size = quadrat_size)
plot_quadrats(domain, q_systematic, "Systematic placement (regular lattice)")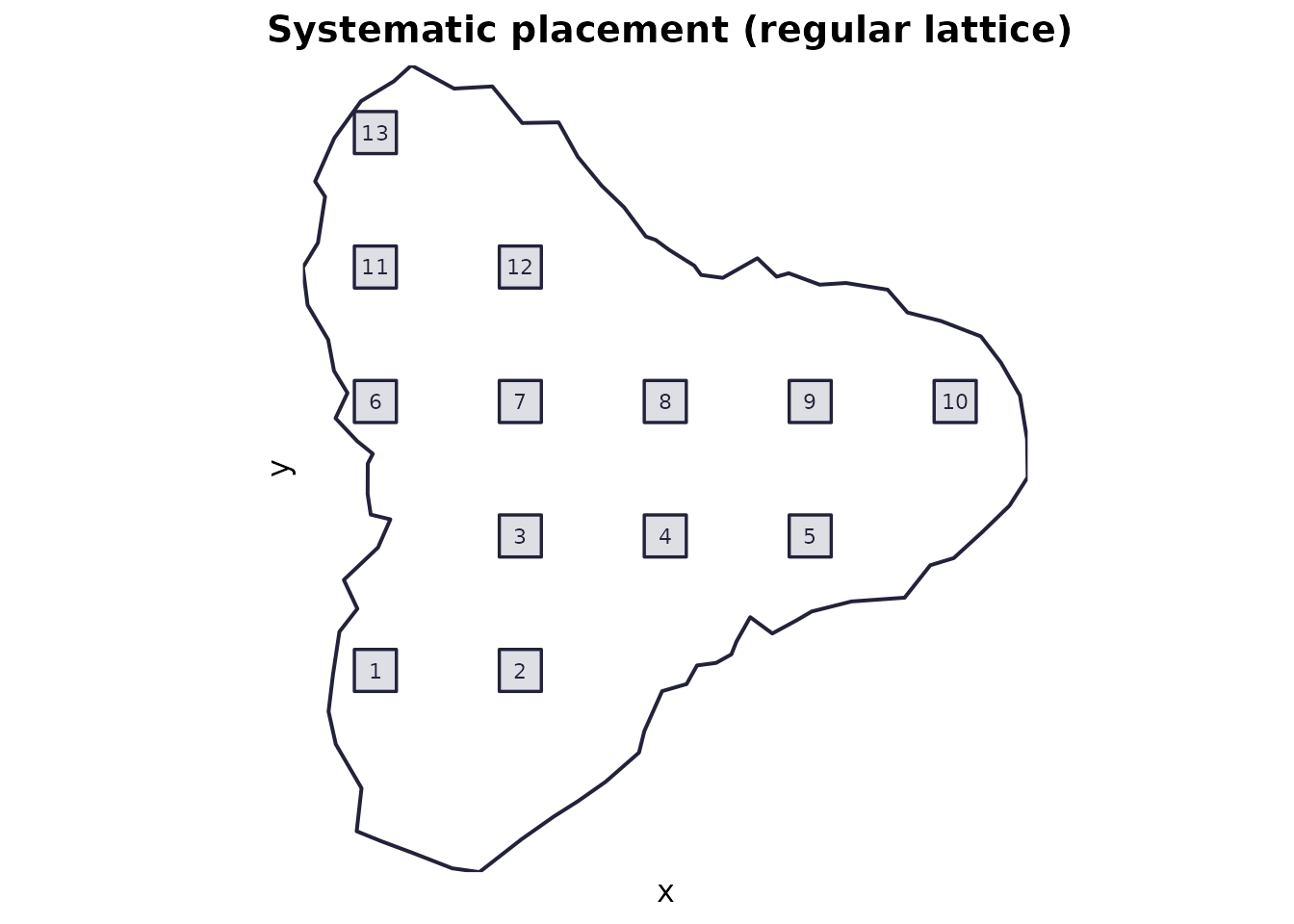
Notes:
- The actual number returned may differ from
n_quadratsbecause the grid is filtered by the domain boundary. - Quadrats may overlap if the grid spacing is smaller than the quadrat size (this depends on the inferred grid density).
4) Transect placement
Use when: you want transect-based sampling (parallel lines across the domain) with evenly spaced quadrats along each transect.
Mechanics: n_transects parallel lines
are created across a safe area (domain shrunk by half the
quadrat diagonal). Along each transect segment,
n_quadrats_per_transect quadrats are placed.
set.seed(5)
q_transect <- place_quadrats_transect(
domain,
n_transects = 2,
n_quadrats_per_transect = 8,
quadrat_size = quadrat_size,
angle = 60
)
plot_quadrats(domain, q_transect, "Transect placement (2 transects, angle = 60°)")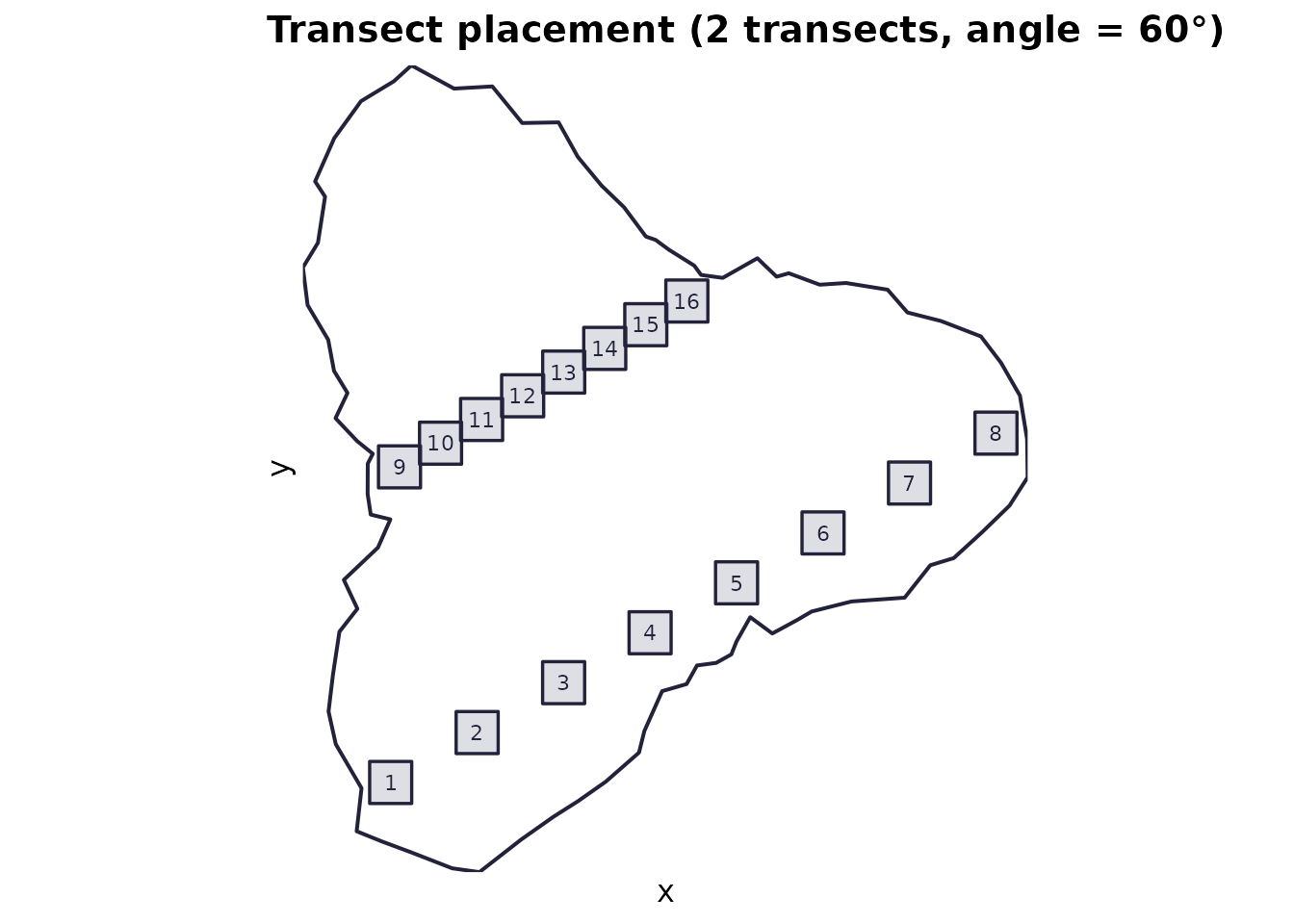
Notes:
- The quadrats themselves remain axis-aligned, even if transects run at an angle.
- If the safe area becomes too small (large quadrats, narrow domain), few or no quadrats may be returned.
5) Voronoi-based placement
Use when: you want sampling locations that are well-spaced in complex domains.
Mechanics: a dense set of random seed points are placed in the domain; a Voronoi tessellation is computed; cells are screened for whether they can contain the quadrat (based on an inscribed circle heuristic). Quadrats are placed at cell centres.
set.seed(6)
# Note: very large voronoi_seed_factor values tend to create many small cells,
# which can make it impossible to fit larger quadrats. If you get an empty
# result, either reduce `voronoi_seed_factor` or use a smaller quadrat_size.
q_voronoi <- place_quadrats_voronoi(
domain,
n_quadrats = 20,
quadrat_size = c(1.2, 1.2),
voronoi_seed_factor = 4,
show_voronoi = TRUE
)
# Standard view (domain + quadrats)
plot_quadrats(domain, q_voronoi, "Voronoi placement (seed_factor = 4; size = 1.2×1.2)")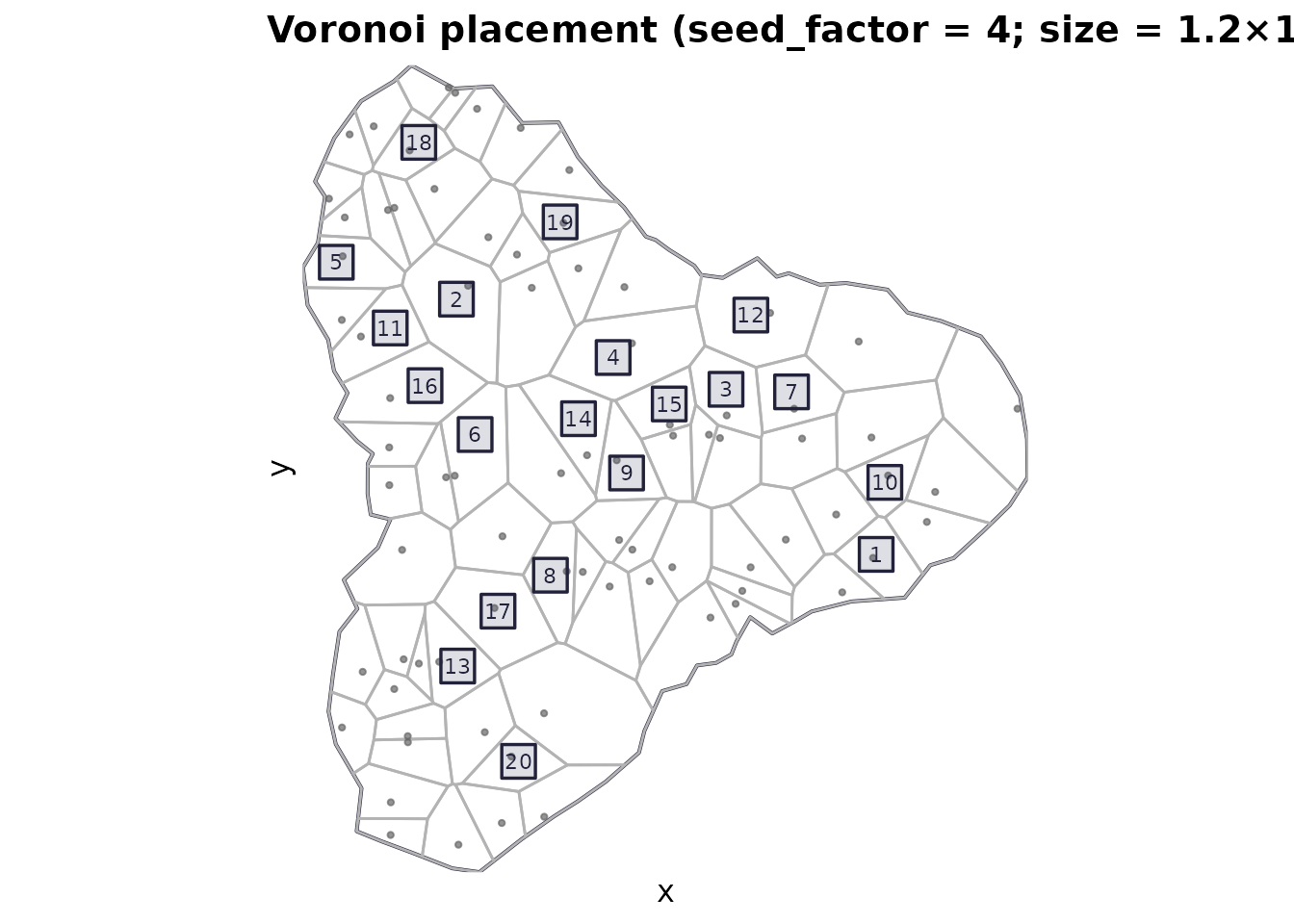
# Optional: show the underlying Voronoi tessellation used to select candidates
vor_cells <- attr(q_voronoi, "voronoi_cells")
seeds <- attr(q_voronoi, "voronoi_seeds")
p_voronoi_diag <- ggplot() +
geom_sf(data = domain, fill = NA, colour = "#22223b", linewidth = 0.7) +
geom_sf(data = vor_cells, fill = NA, colour = "grey65", linewidth = 0.5) +
geom_sf(data = sf::st_as_sf(seeds), colour = "grey40", alpha = 0.7, size = 0.8) +
geom_sf(data = q_voronoi, fill = "#4a4e69", alpha = 0.18, colour = "#22223b", linewidth = 0.6) +
coord_sf(datum = NA) +
labs(title = "Voronoi diagram (clipped) + seeds + selected quadrats") +
theme_minimal(base_size = 12) +
theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank())
print(p_voronoi_diag)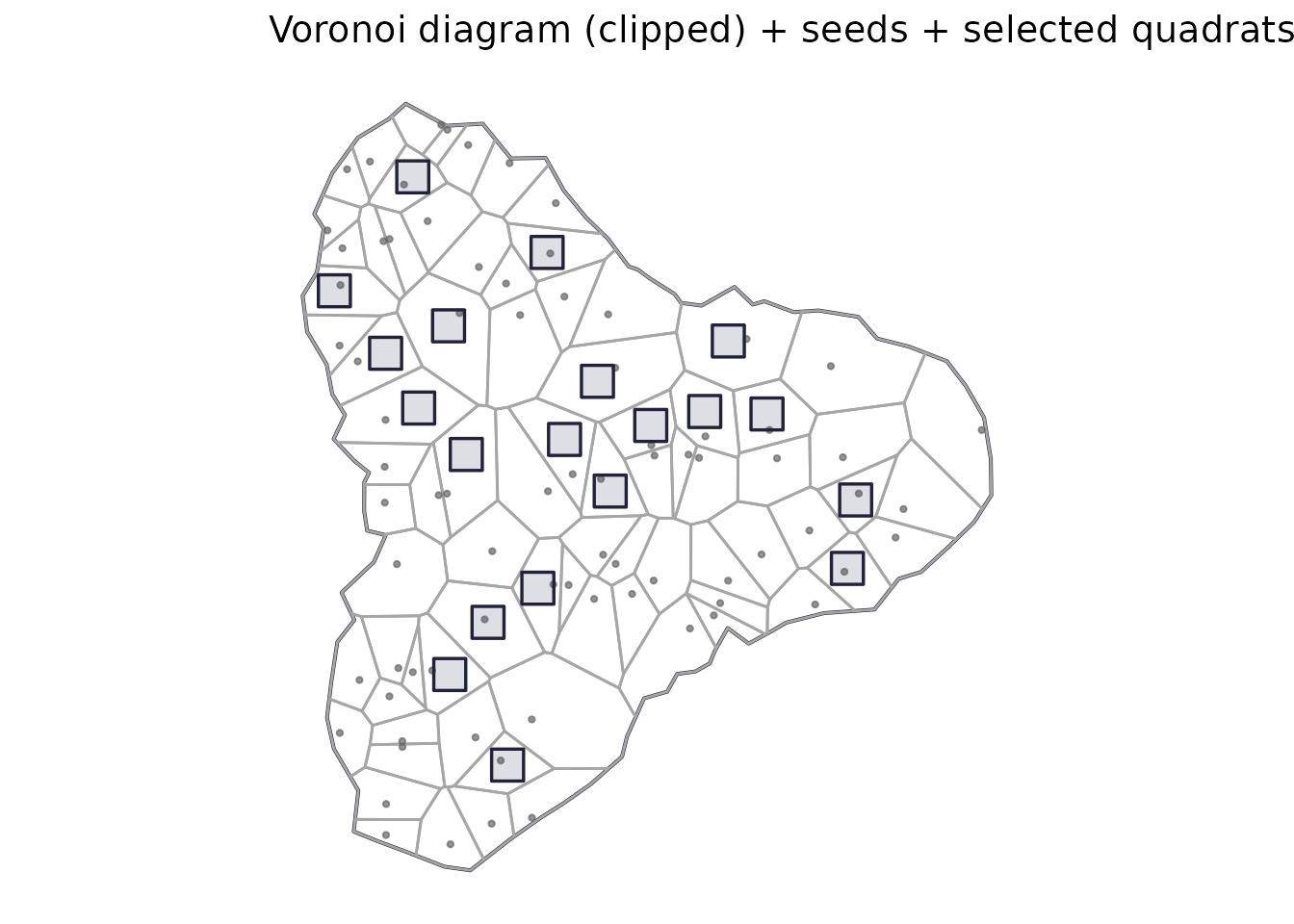
Notes:
- For narrow or very irregular domains, increase
voronoi_seed_factor(at a computational cost). - The current implementation focuses on finding feasible, nicely spaced sites; it is not designed to guarantee strict non-overlap in all cases (though overlap is uncommon for moderate sizes).
Summary: when to use which
- Random: general-purpose, classic random sampling, non-overlapping.
- Tiled: systematic grid cells; great for “choose random cells from a tiling”.
- Systematic: classic lattice-based systematic sampling.
- Transect: teaching/field designs with transects.
- Voronoi: irregular domains where you want good spatial coverage and spacing.
Reproducibility tip
All schemes are stochastic to some degree (even the systematic scheme can depend on inferred grid dimensions); for reproducible figures, set a seed immediately before each placement call.
Method-testing tip: inspect placement audits (boundary exclusion)
Quadrat placement can fail silently in subtle ways when the domain is irregular (e.g., many candidates fall outside the boundary, or random placement hits an attempt cap).
spesim attaches a placement_audit attribute to the
returned quadrats. This is used by
spesim_audit()/audit_sampling_scheme(), and it
is also useful to inspect directly.
library(spesim)
set.seed(1)
dom <- create_sampling_domain()
q_rand <- place_quadrats(dom, n_quadrats = 20, quadrat_size = c(1.5, 1.5))
attr(q_rand, "placement_audit")
q_tile <- place_quadrats_tiled(dom, n_quadrats = 20, quadrat_size = c(1.5, 1.5))
attr(q_tile, "placement_audit")